Factors to Death
Data Importing
source("data_preprocessing_backbone.R")
labor_data = read_csv("./data/CA_Labor.csv") %>%
janitor::clean_names() %>%
dplyr::select(-employment,-unemployment,-rank) %>%
rename(unemployment=unemployment_rate_per_cent)
land_data = read_csv("./data/CA_Land_Area.csv") %>%
janitor::clean_names() %>%
mutate(
location = ifelse(county %in% c("Los Angeles", "Orange", "San Diego", "Monterey", "San Benito", "San Luis Obispo", "Santa Barbara", "Santa Cruz", "Ventura", "Alameda", "Contra Costa", "Marin", "Napa", "San Francisco", "San Mateo", "Santa Clara", "Solano", "Sonoma", "Del Norte", "Humboldt", "Mendocino"), "costal", "inland")) %>%
dplyr::select(-rank, -state, -population)
ca_nonmedical_data = left_join(land_data, labor_data, by = c("county"))
demo_covid = demo %>%
mutate(cumulative_reported_deaths= as.numeric(cumulative_reported_deaths),
cumulative_cases = as.numeric(cumulative_cases),
death_rate = 0.01+cumulative_reported_deaths/population*100,
test = cumulative_total_tests/population*100) %>%
dplyr::select(county_name, death_rate, test,population) %>%
rename(county=county_name) %>%
group_by(county) %>%
summarize(death_rate=max(death_rate,na.rm = T),
test = max(test),
population = max(population)) %>%
filter(!county %in% c("Out of state","Unknown"))
vaccine = read_csv("./data/CA_covid19vaccines.csv") %>%
janitor::clean_names() %>%
group_by(county) %>%
summarise(
fully_vaccinated = sum(fully_vaccinated)
# total number of fully vaccinated people
) %>%
arrange(county) %>%
drop_na() %>%
filter(!county %in% c("Unknown", "All CA Counties", "All CA and Non-CA Counties","Outside California"))
hospital = read_csv("./data/CA_Covid19_Hospital.csv") %>%
group_by(county) %>%
summarize(all_hospital_beds = max(all_hospital_beds,na.rm = T),
icu_available_beds = max(icu_available_beds,na.rm = T)) %>%
dplyr::select(county, all_hospital_beds, icu_available_beds)
ca_medical_data = left_join(demo_covid, vaccine, by =c("county"))
ca_medical_data2 = left_join(ca_medical_data, hospital, by =c("county"))
ca_premodel_data = left_join(ca_medical_data2, ca_nonmedical_data, by = c("county")) %>%
relocate(death_rate) %>%
mutate(
vaccinated=fully_vaccinated/population*100,
labor_rate = labor_force/population*100,
hospital_bed = all_hospital_beds/population*100,
icu_bed = icu_available_beds/population*100
)%>%
dplyr::select(-fully_vaccinated, -population, -labor_force,-all_hospital_beds,-icu_available_beds)
skimr::skim(ca_premodel_data)| Name | ca_premodel_data |
| Number of rows | 58 |
| Number of columns | 10 |
| _______________________ | |
| Column type frequency: | |
| character | 2 |
| numeric | 8 |
| ________________________ | |
| Group variables | None |
Variable type: character
| skim_variable | n_missing | complete_rate | min | max | empty | n_unique | whitespace |
|---|---|---|---|---|---|---|---|
| county | 0 | 1 | 4 | 15 | 0 | 58 | 0 |
| location | 0 | 1 | 6 | 6 | 0 | 2 | 0 |
Variable type: numeric
| skim_variable | n_missing | complete_rate | mean | sd | p0 | p25 | p50 | p75 | p100 | hist |
|---|---|---|---|---|---|---|---|---|---|---|
| death_rate | 0 | 1.00 | 0.21 | 0.09 | 0.01 | 0.13 | 0.19 | 0.28 | 0.51 | ▂▇▅▃▁ |
| test | 0 | 1.00 | 367.42 | 171.84 | 104.51 | 277.33 | 312.06 | 402.06 | 983.95 | ▅▇▂▁▁ |
| land_area_sq_mi | 0 | 1.00 | 2685.85 | 3102.32 | 46.87 | 959.49 | 1535.34 | 3454.40 | 20056.92 | ▇▂▁▁▁ |
| unemployment | 0 | 1.00 | 7.33 | 2.12 | 4.42 | 5.97 | 6.96 | 8.22 | 17.37 | ▇▆▁▁▁ |
| vaccinated | 0 | 1.00 | 63.20 | 13.47 | 29.78 | 54.34 | 61.29 | 72.35 | 94.09 | ▁▇▇▇▃ |
| labor_rate | 0 | 1.00 | 44.16 | 5.95 | 30.47 | 40.29 | 44.59 | 47.75 | 61.19 | ▂▅▇▁▁ |
| hospital_bed | 2 | 0.97 | 0.32 | 0.39 | 0.08 | 0.18 | 0.22 | 0.29 | 2.79 | ▇▁▁▁▁ |
| icu_bed | 2 | 0.97 | 0.03 | 0.05 | 0.00 | 0.01 | 0.01 | 0.03 | 0.35 | ▇▁▁▁▁ |
- The data includes 58 observations and 10 variables. Now we are
interested on whether the death cases are associated with
area,populationandtesting_cases.
Background
- We were curious about what medical and non-medical factors were associated with death rates, so we collected labor force data, population data as non-medical data for each county in California, using the percentage value of the largest labor force to population over a three-year range as an important indicator of local economic conditions, in addition to unemployment rate data, which is the ratio of the unemployed population divided by the labor force. Also, we collected COVID infection death data, vaccination data from each county, hospital data and defined the mortality rate as \(\frac{number\ of\ deaths}{maximum\ total\ population}\) over 2020-2022, and the vaccination rate as \(\frac{fully\ vaccinated\ population}{maximum\ total\ population}\) over the three-year period.
Data processing
- Labor force data: Because total population data are not available in this dataset, calculating the labor force rate requires demographic variables from the population dataset. Therefore, I extracted the county name, unemployment rate, and labor force variables.
- Demographic data: We extracted county area, population, and divided the counties into coastal and inland counties based on the official definition of coastal counties as another important potential economic indicator.
- Labor force data and total population data were combined by
left_joinand divided by labor force by population as labor force percentage - COVID data: daily covid infection death data for each county, I divided the total number of infections by the population as the death rate and the total number of tests by the population as the nucleic acid detection rate, and finally the county was used as the county, and the largest value over three years was taken as the indicator for that county.
- Vaccine data: Similar to COVID data, the number of full vaccination completions divided by the total population was extracted as the vaccination rate, and the largest value over three years was taken as the indicator for that county.
- Hospital data: The maximum number of beds and the maximum number of available ICU beds in each county over three years were taken as two potentially important medical indicators.
- The above data were combined to obtain pre-modeling pre-modelling
data, including death_rate, county, test, land_area_sq_mi, location,
unemployment, vaccinated, labor_rate, hospital_bed, icu_bed. This
pre-modelling data includes 58 observations and 10 variables, and two
cells in each
hospital_bed,icu_bed. Now we are interested on whether the death cases are associated with these selected variables.
Data Description
point <- format_format(big.mark = " ", decimal.mark = ",", scientific = FALSE)
raw_deaths = ca_premodel_data %>%
ggplot(aes(x = death_rate)) +
geom_density(fill = "#69b3a2", color = "#e9ecef", alpha = .8) +
scale_y_continuous(labels = point)
raw_tests = ca_premodel_data %>%
ggplot(aes(x = test)) +
geom_density(fill = "#69b3a2", color = "#e9ecef", alpha = .8) +
scale_y_continuous(labels = point) +
scale_x_continuous(labels = point)
raw_land = ca_premodel_data %>%
ggplot(aes(x = land_area_sq_mi)) +
geom_density(fill = "#69b3a2", color = "#e9ecef", alpha = .8) +
scale_y_continuous(labels = point) +
scale_x_continuous(labels = point)
raw_unemployment= ca_premodel_data %>%
ggplot(aes(x = unemployment)) +
geom_density(fill = "#69b3a2", color = "#e9ecef", alpha = .8) +
scale_y_continuous(labels = point) +
scale_x_continuous(labels = point)
raw_vaccinated = ca_premodel_data %>%
ggplot(aes(x = vaccinated)) +
geom_density(fill = "#69b3a2", color = "#e9ecef", alpha = .8)
raw_labor = ca_premodel_data %>%
ggplot(aes(x = labor_rate)) +
geom_density(fill = "#69b3a2", color = "#e9ecef", alpha = .8)
raw_hospital = ca_premodel_data %>%
ggplot(aes(x = hospital_bed)) +
geom_density(fill = "#69b3a2", color = "#e9ecef", alpha = .8)
raw_bed = ca_premodel_data %>%
ggplot(aes(x = icu_bed)) +
geom_density(fill = "#69b3a2", color = "#e9ecef", alpha = .8)
ca_model_data = ca_premodel_data %>%
mutate(ln_tests = log(test),
ln_area = log(land_area_sq_mi),
ln_unemployment = log(unemployment),
ln_hospital = log(hospital_bed+0.01),
ln_bed = log(icu_bed+0.01)
) %>%
dplyr::select(-test,-land_area_sq_mi,-unemployment,-county,-hospital_bed,-icu_bed) %>%
filter(!ln_hospital %in% NA,
!ln_bed %in% NA)
plot_ln_test = ca_model_data %>%
ggplot(aes(x = ln_tests)) +
geom_density(fill = "#69b3a2", color = "#e9ecef", alpha = .8)
plot_ln_hospital = ca_model_data %>%
ggplot(aes(x = ln_hospital)) +
geom_density(fill = "#69b3a2", color = "#e9ecef", alpha = .8)
plot_ln_bed = ca_model_data %>%
ggplot(aes(x = ln_bed)) +
geom_density(fill = "#69b3a2", color = "#e9ecef", alpha = .8)
plot_ln_area = ca_model_data %>%
ggplot(aes(x = ln_area)) +
geom_density(fill = "#69b3a2", color = "#e9ecef", alpha = .8)
plot_ln_unemployment = ca_model_data %>%
ggplot(aes(x = ln_unemployment)) +
geom_density(fill = "#69b3a2", color = "#e9ecef", alpha = .8)
raw_deaths+raw_tests+raw_land+ raw_unemployment+raw_hospital +raw_bed+ raw_vaccinated+ raw_labor+ plot_ln_test+ plot_ln_area+ plot_ln_unemployment+plot_ln_hospital+plot_ln_bed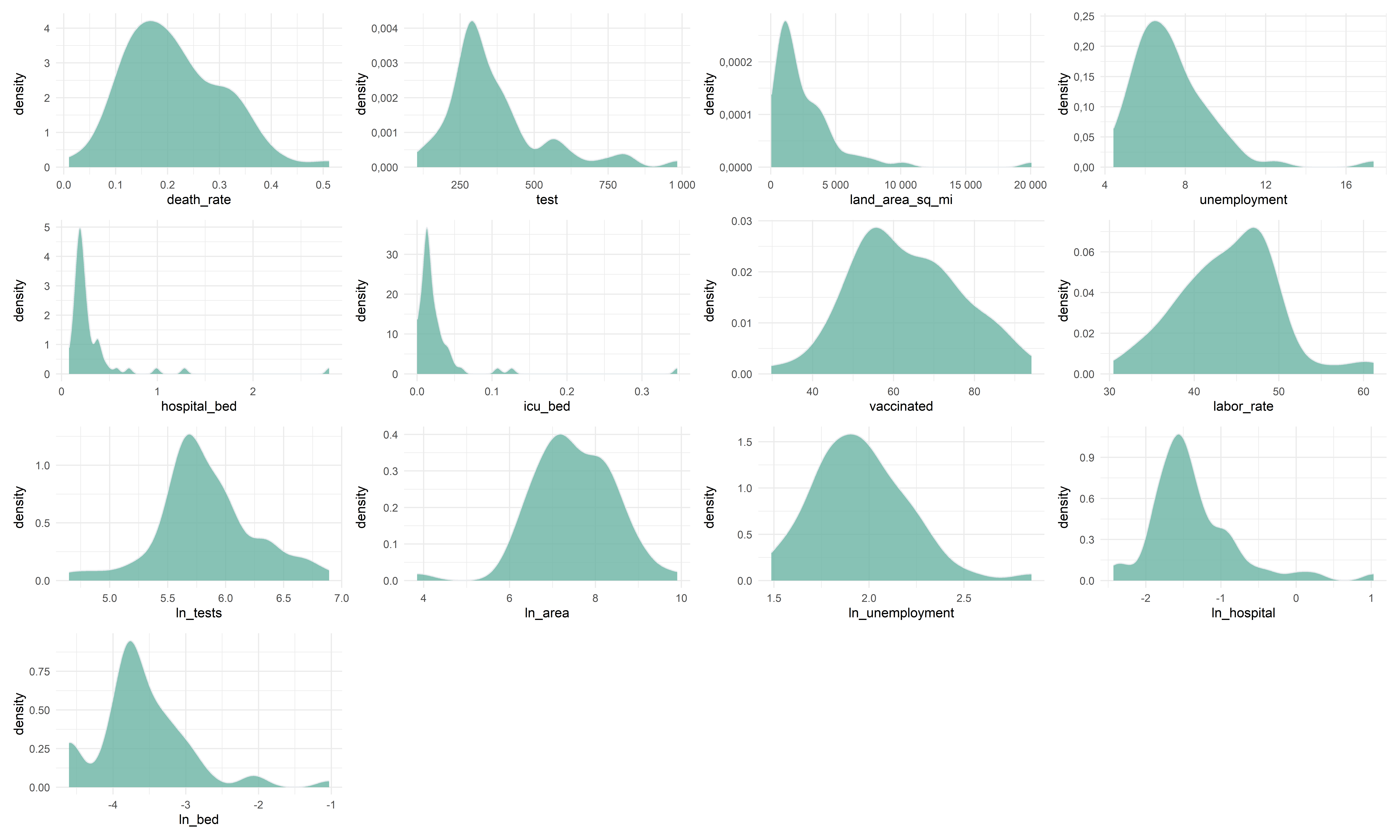
ca_model_data %>%
gtsummary::tbl_summary() %>%
gtsummary::bold_labels() %>%
as.tibble() %>%
knitr::kable()| Characteristic | N = 56 |
|---|---|
| death_rate | 0.20 (0.14, 0.28) |
| location | NA |
| costal | 21 (38%) |
| inland | 35 (62%) |
| vaccinated | 63 (54, 73) |
| labor_rate | 44.6 (39.9, 47.8) |
| ln_tests | 5.75 (5.63, 6.02) |
| ln_area | 7.38 (6.90, 8.17) |
| ln_unemployment | 1.94 (1.78, 2.11) |
| ln_hospital | -1.49 (-1.68, -1.21) |
| ln_bed | -3.71 (-3.87, -3.30) |
- The cleaned data includes 56 states and 9 variables including death_rate, location, vaccinated, labor_rate, ln_tests, ln_area, ln_unemployment, ln_hospital, ln_bed. For those variables whose data is extremely right-skewed and the natural logarithms are applied to each exploratory continuous variables. By the plots and summary table we can see all the variables are almost bell-shaped, so in the modelling dataset we deleted the original variables that need to be transformed, keep the transformed variables, and delete the county variable because the county variable itself does not play a role in the model.
Correlation
ca_model_data %>%
dplyr::select(-location) %>%
GGally::ggpairs(upper = list(continuous = GGally::wrap("cor", size = 3)))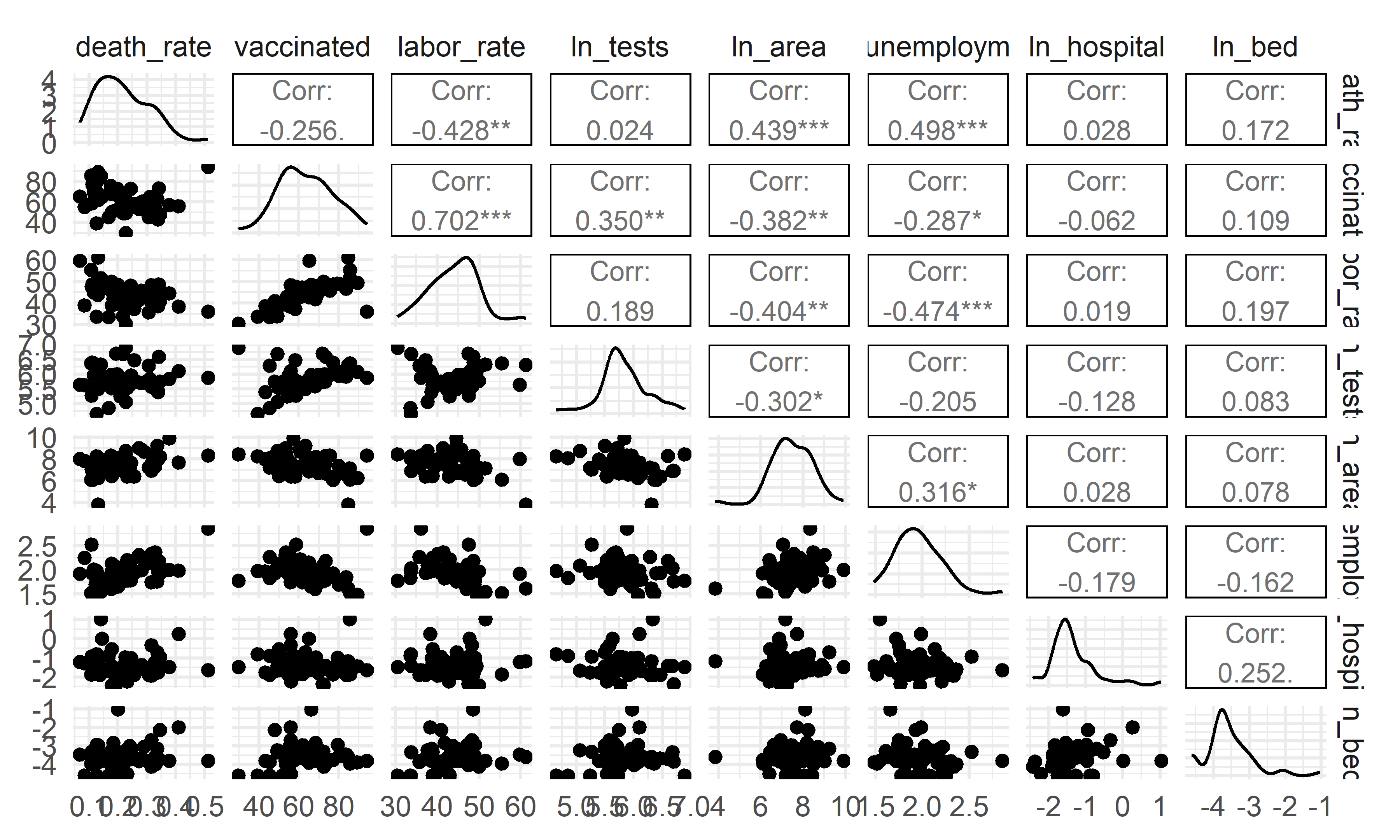
- The overall correlation and covariance between the independent
variables are not obvious, but among them we can see that there is a
large trend of correlation and covariance between
vaccinationandlabor_rate, implying that including that including both of these variables in the model could result in poor coefficient estimation and inflated standard errors (due to multicollinearity). Our model selection techniques are able to figure this out.
Modelling Fit
reg_full = lm(death_rate ~ ., data = ca_model_data)
reg_intercept = lm(death_rate ~ 1, data = ca_model_data)- We fitted a model with all potential predictors and a model including only the intercept for proceeding automatic model selection.
Model Selection
#forward Selection
step(reg_intercept, direction='forward', scope=formula(reg_full),trace=0) %>%
broom::tidy() %>%
mutate(term = str_replace(term, "^location", "Location: ")) %>%
knitr::kable(digits = 3)| term | estimate | std.error | statistic | p.value |
|---|---|---|---|---|
| (Intercept) | -0.560 | 0.193 | -2.899 | 0.006 |
| ln_unemployment | 0.127 | 0.041 | 3.059 | 0.004 |
| ln_area | 0.028 | 0.010 | 2.637 | 0.011 |
| ln_bed | 0.032 | 0.014 | 2.212 | 0.032 |
| Location: inland | 0.067 | 0.023 | 2.946 | 0.005 |
| ln_tests | 0.068 | 0.024 | 2.818 | 0.007 |
#backwards elimination
step(reg_full, direction='backward', scope=formula(reg_full),trace=0) %>%
broom::tidy() %>%
mutate(term = str_replace(term, "^location", "Location: ")) %>%
knitr::kable(digits = 3)| term | estimate | std.error | statistic | p.value |
|---|---|---|---|---|
| (Intercept) | -0.375 | 0.245 | -1.528 | 0.133 |
| Location: inland | 0.086 | 0.027 | 3.136 | 0.003 |
| vaccinated | 0.002 | 0.001 | 1.828 | 0.074 |
| labor_rate | -0.004 | 0.002 | -1.878 | 0.066 |
| ln_tests | 0.060 | 0.024 | 2.510 | 0.016 |
| ln_area | 0.026 | 0.011 | 2.438 | 0.019 |
| ln_unemployment | 0.093 | 0.044 | 2.139 | 0.038 |
| ln_bed | 0.035 | 0.014 | 2.479 | 0.017 |
#stepwise Selection
step(reg_full, direction = "both", scope=formula(reg_full),trace=0) %>%
broom::tidy() %>%
mutate(term = str_replace(term, "^location", "Location: ")) %>%
knitr::kable(digits = 3)| term | estimate | std.error | statistic | p.value |
|---|---|---|---|---|
| (Intercept) | -0.375 | 0.245 | -1.528 | 0.133 |
| Location: inland | 0.086 | 0.027 | 3.136 | 0.003 |
| vaccinated | 0.002 | 0.001 | 1.828 | 0.074 |
| labor_rate | -0.004 | 0.002 | -1.878 | 0.066 |
| ln_tests | 0.060 | 0.024 | 2.510 | 0.016 |
| ln_area | 0.026 | 0.011 | 2.438 | 0.019 |
| ln_unemployment | 0.093 | 0.044 | 2.139 | 0.038 |
| ln_bed | 0.035 | 0.014 | 2.479 | 0.017 |
# Use criterion-based procedures to guide your selection of the ‘best subset’
# chosen using SSE/RSS (smaller is better)
criterion_selected = regsubsets(death_rate ~ ., data = ca_model_data)
criterion_plot = summary(criterion_selected)
# plot of Cp and Adj-R2 as functions of parameters
x_fe = (2:9)
y_fe = criterion_plot$cp
z_fe = criterion_plot$adjr2
cp_fe_p =
cbind(x_fe, y_fe, z_fe) %>%
as.tibble() %>%
rename(`Number of parameter` = x_fe,
`Cp value` = y_fe,
`Adj R-square` = z_fe) %>%
mutate(p = `Number of parameter`)
cp_fe_p_1 =
cp_fe_p %>%
ggplot() +
geom_point(aes(x = `Number of parameter`, y = `Cp value`, color = `Cp value`)) +
theme(legend.position = "none") +
geom_abline(intercept = 0, slope = 1, color = "red")
cp_fe_p_2 =
cp_fe_p %>%
ggplot() +
geom_point(aes(x = `Number of parameter`, y = `Adj R-square`, color = `Adj R-square`)) +
theme(legend.position = "none")
cp_fe_p_1 + cp_fe_p_2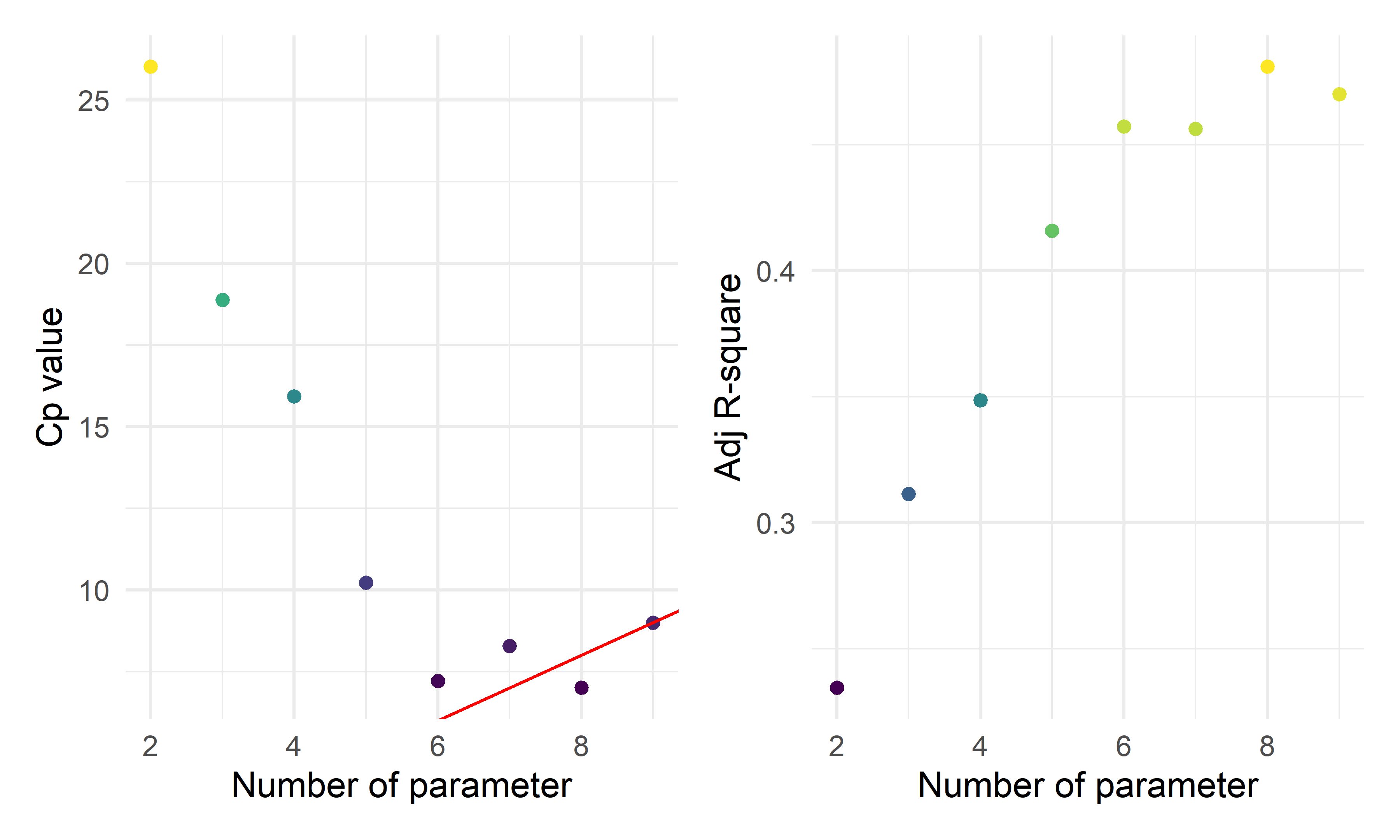
- By the two automatic model selection models: backward elimination and stepwise selection, we got the same result containing eight parameters (seven predictors):
\(\hat{death\_rate}=-0.375+0.086*I(location=inland)+0.002*vac-0.004*labor+0.060*ln(tests)+0.026*ln(area)+0.093*ln(unemploy)+0.035*ln(bed)\) where vac is the vaccination rate, labor is the labor rate, unemploy is the unemployment rate, bed is the hospital available beds per person.
However, forward selection deleted
vaccinatedandlabor_rate, whose p-values also exceed 0.05 in the other two models.The plots above show the \(C_p\) index and Adjusted \(R^2\) for various numbers of parameters. When choosing a model based on \(C_p\) criterion, we want to choose a model for which \(C_p \leq p\), where p is the number of parameters. From the \(C_p\) plot above, we should have 8 parameters (7 predictors), it has the lowest value for \(C_p\) compared with the number of parameters and the highest \(R^2\).
Penalized Model Selection Method: LASSO
ca_lasso_data = ca_model_data %>% mutate(
location=ifelse(location == "costal", 1,0))
# using cross validation to choose lambda
lambda_seq = 10^seq(-5, -1, by = .01)
set.seed(2022)
cv_object <- cv.glmnet(as.matrix(ca_lasso_data[2:9]), ca_lasso_data$death_rate,
lambda = lambda_seq, nfolds = 5)
cv_object##
## Call: cv.glmnet(x = as.matrix(ca_lasso_data[2:9]), y = ca_lasso_data$death_rate, lambda = lambda_seq, nfolds = 5)
##
## Measure: Mean-Squared Error
##
## Lambda Index Measure SE Nonzero
## min 0.002692 158 0.006254 0.001357 7
## 1se 0.019498 72 0.007608 0.001773 4# plot the CV results
tibble(lambda = cv_object$lambda,
mean_cv_error = cv_object$cvm) %>%
mutate(text_label = str_c("Lambda: ", lambda,
"\n Mean CV error: ", mean_cv_error)) %>%
plotly::plot_ly(y = ~mean_cv_error, x = ~lambda,
color = ~lambda,
width = 900,
height = 500,
type = "scatter",
mode = "markers",
marker = list(size = 6),
colors = "magma",
text = ~ text_label)# refit the lasso model with the "best" lambda
fit_bestcv <- glmnet(as.matrix(ca_lasso_data[2:9]), ca_model_data$death_rate, lambda = cv_object$lambda.min)
fit_bestcv %>%
broom::tidy() %>%
mutate(term = str_replace(term, "^location", "Location: ")) %>%
knitr::kable(digits = 3)| term | step | estimate | lambda | dev.ratio |
|---|---|---|---|---|
| (Intercept) | 1 | -0.251 | 0.003 | 0.534 |
| Location: | 1 | -0.066 | 0.003 | 0.534 |
| vaccinated | 1 | 0.001 | 0.003 | 0.534 |
| labor_rate | 1 | -0.003 | 0.003 | 0.534 |
| ln_tests | 1 | 0.051 | 0.003 | 0.534 |
| ln_area | 1 | 0.023 | 0.003 | 0.534 |
| ln_unemployment | 1 | 0.099 | 0.003 | 0.534 |
| ln_bed | 1 | 0.030 | 0.003 | 0.534 |
To make it easier for LASSO to proceed, I converted the
positionvariables to 0 and 1. Using the LASSO method to perform variable selection, the key step is to make sure choosing the optimal \(\lambda\) to efficiently reduce the complexity of the model while some descriptive prediction ability maintained. Thus a five-fold cross validation was performed.Since the two stepwise selection techniques and the criterion techniques all chose the same model with 7 predictors, we recommend this as our final model. Also, the LASSO gave a very similar suggested model.
Fitting the seleted final model
mlr_model = lm(death_rate~location+vaccinated+labor_rate+ln_tests+ln_area+ln_unemployment+ln_bed,data = ca_model_data)
mlr_model %>%
broom::tidy() %>%
mutate(term = str_replace(term, "^location", "Location: ")) %>%
knitr::kable(digits = 3)| term | estimate | std.error | statistic | p.value |
|---|---|---|---|---|
| (Intercept) | -0.375 | 0.245 | -1.528 | 0.133 |
| Location: inland | 0.086 | 0.027 | 3.136 | 0.003 |
| vaccinated | 0.002 | 0.001 | 1.828 | 0.074 |
| labor_rate | -0.004 | 0.002 | -1.878 | 0.066 |
| ln_tests | 0.060 | 0.024 | 2.510 | 0.016 |
| ln_area | 0.026 | 0.011 | 2.438 | 0.019 |
| ln_unemployment | 0.093 | 0.044 | 2.139 | 0.038 |
| ln_bed | 0.035 | 0.014 | 2.479 | 0.017 |
As almost all the terms has p-value < 0.05, with the
locationvariable < 0.1, we can conclude that all the predictor coefficients are statistical significant at 0.1 significance level and 0.05 significance level forlocation,ln_test,ln_area,ln_unemployment,ln_bed. Hence all the seven variables will be included in the model. All these methods almost agree on the same suggested model or predictors. This might give good evidence that we should explore this model further.From the outputs above, we see that
vaccinatedandlabor_ratehad a p-value more than 0.05. This might mean that the relationship betweendeath_rateand these two variables is weak. Also, the highly correlated variablesvaccinatedandlabor_ratewere both included in the final models. The possible reason for this is that the model is slightly underfitted and there are potential predictors that are not included in the dataset.
Modelling Diagnosis
check_model(mlr_model, check = c("linearity", "qq", "normality", "outliers", "homogeneity", "vif"))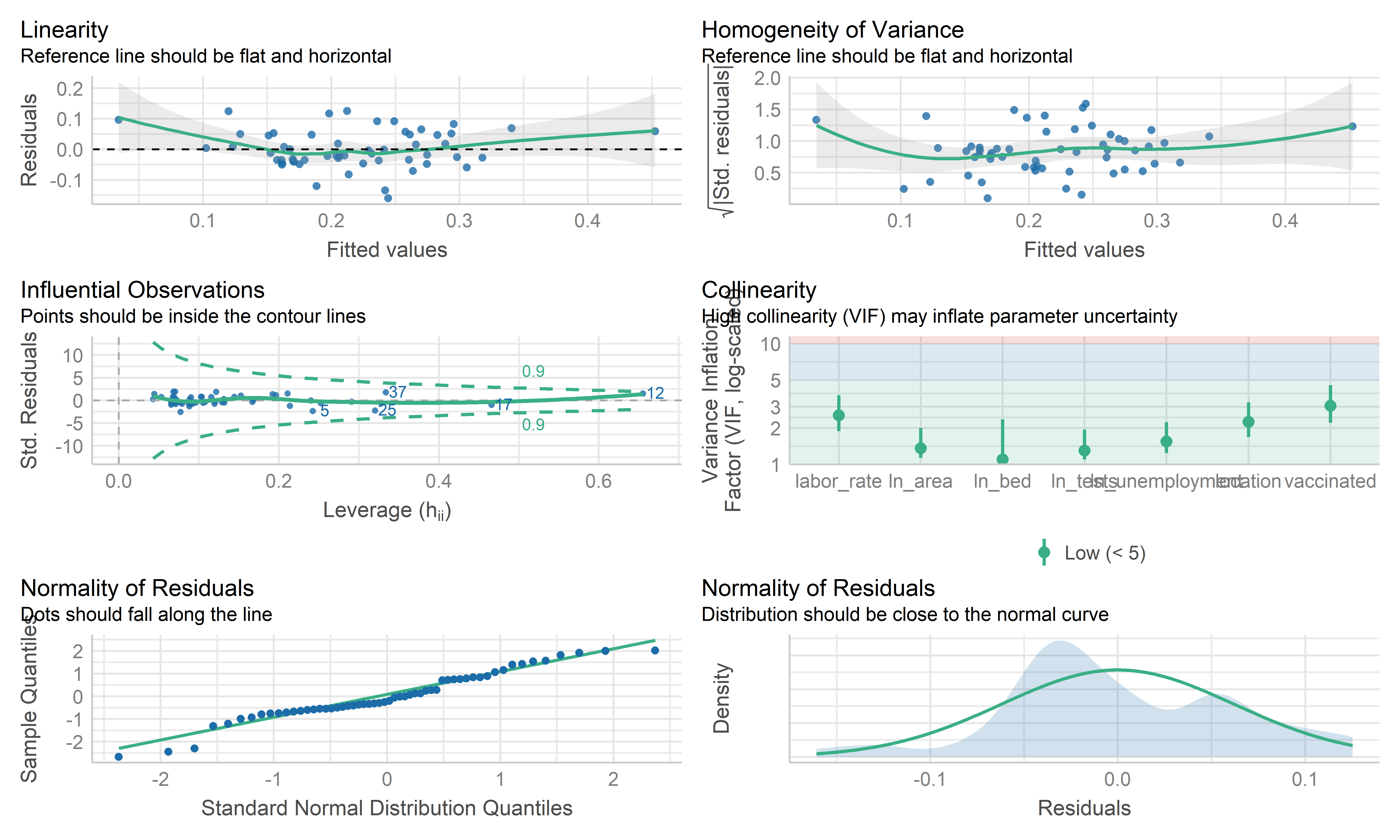
- Overall, these plots look like this model is fitting the data well. No influential point detected. However the residual points showed some kinds trend and the horizontal line is not such stable on the edge area, it may be worth investigating further by weighted least squares regression.
Weighted Least Squares Regression
# define sd function
sd_function <- lm(abs(reg_full$residuals) ~ reg_full$fitted.values)
var_fitted <- sd_function$fitted.values^2
#define weight
wt <- 1/var_fitted
wls_full = lm(death_rate~.,data = ca_model_data, weights = wt)
wls_death = step(wls_full, direction='backward', scope=formula(wls_full),trace=0)
wls_death %>%
broom::tidy() %>%
mutate(term = str_replace(term, "^location", "Location: ")) %>%
knitr::kable(digits = 3)| term | estimate | std.error | statistic | p.value |
|---|---|---|---|---|
| (Intercept) | -0.357 | 0.247 | -1.447 | 0.154 |
| Location: inland | 0.083 | 0.028 | 3.010 | 0.004 |
| vaccinated | 0.002 | 0.001 | 1.557 | 0.126 |
| labor_rate | -0.004 | 0.002 | -1.718 | 0.092 |
| ln_tests | 0.059 | 0.024 | 2.468 | 0.017 |
| ln_area | 0.024 | 0.011 | 2.255 | 0.029 |
| ln_unemployment | 0.093 | 0.044 | 2.103 | 0.041 |
| ln_bed | 0.034 | 0.014 | 2.373 | 0.022 |
check_model(wls_death, check = c("linearity", "qq", "normality", "outliers", "homogeneity", "vif"))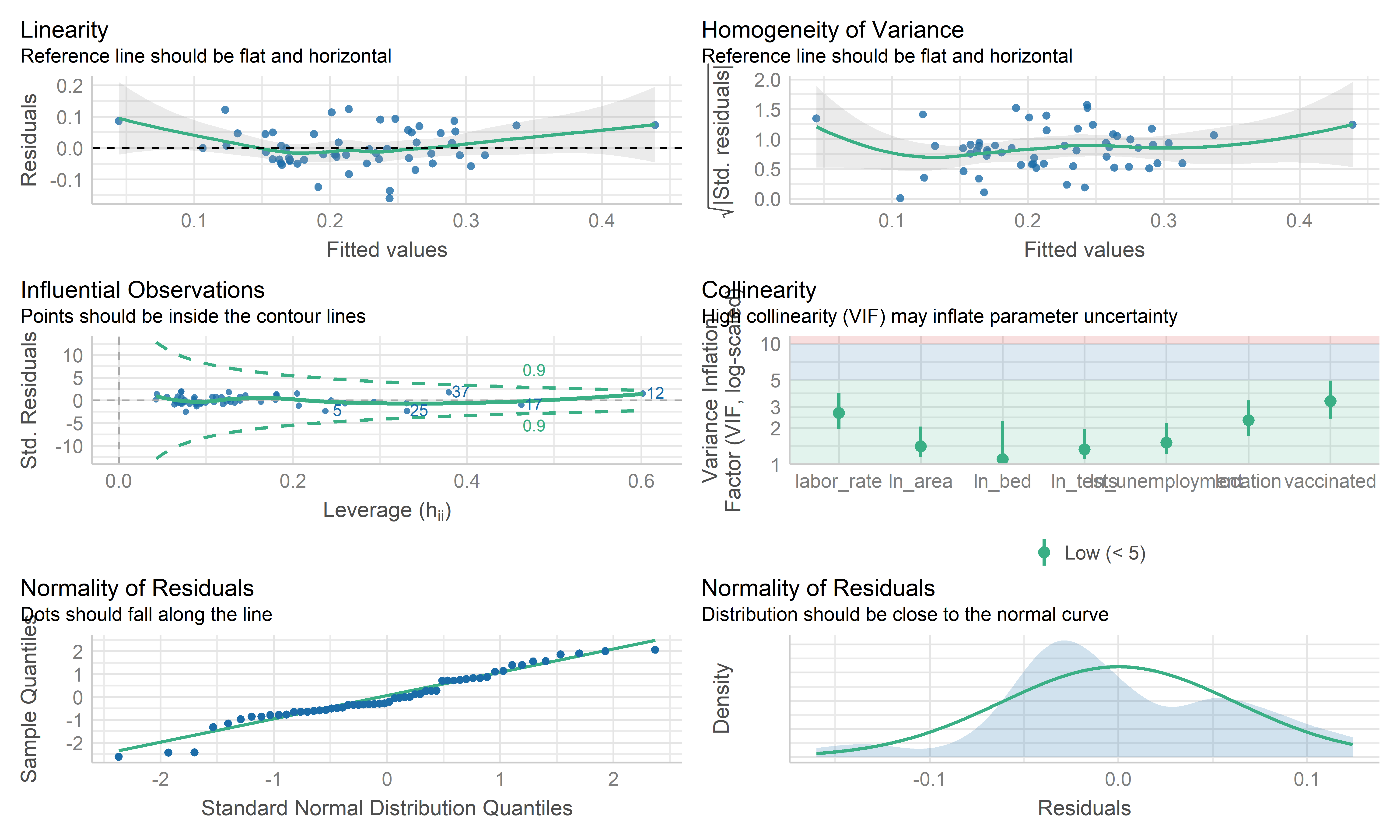
- The coefficient estimate for the predictors variable changed
somewhat and the model fit improved, while
vaccinatedvariable became more insignificant. - The residual standard error per coefficient slightly changed in the weighted LS model. This shows that the predictions from the WLS model are much closer to the actual observations than those from the ordinary LS model.
- \(R^2\) improved in the WLS model. WLS model is able to explain more of the variation in death compared to the OLS model.
- Therefore, I regard both MLR’s mod and WLS’s mod as good models. Because the data are collected from the overall California county-level big data, each observation may contain some special information rather than typo or anything else. This modeling mainly focuses on exploring the predictors used to explain the death, therefore, the MLR model is used here as the final model, while WLS can be used for potential prediction.
Cross Validation
set.seed(2022)
# use 10-fold validation and create the training sets
train = trainControl(method = "cv", number = 10)
# fit the 4-variables model that we selected as our final model
model_caret = train(death_rate~location+vaccinated+labor_rate+ln_tests+ln_area+ln_unemployment+ln_bed, data = ca_model_data, trControl = train, method = 'lm', na.action = na.pass)
model_caret## Linear Regression
##
## 56 samples
## 7 predictor
##
## No pre-processing
## Resampling: Cross-Validated (10 fold)
## Summary of sample sizes: 51, 51, 49, 52, 50, 50, ...
## Resampling results:
##
## RMSE Rsquared MAE
## 0.07404814 0.4120571 0.05966561
##
## Tuning parameter 'intercept' was held constant at a value of TRUEmodel_caret$resample## RMSE Rsquared MAE Resample
## 1 0.04542436 7.158233e-01 0.04036246 Fold01
## 2 0.09564178 3.185436e-05 0.07344651 Fold02
## 3 0.05735306 3.725184e-01 0.05050860 Fold03
## 4 0.07528233 3.078636e-01 0.05997718 Fold04
## 5 0.05641814 5.340497e-01 0.05120324 Fold05
## 6 0.07235722 2.402190e-01 0.06018088 Fold06
## 7 0.08197426 4.776110e-01 0.06190731 Fold07
## 8 0.08782930 2.459130e-01 0.06642902 Fold08
## 9 0.09308560 3.665463e-01 0.07095667 Fold09
## 10 0.07511538 8.599950e-01 0.06168425 Fold10- From the output above, the overall RMSE (root mean squared error) is 0.074, which would mean our MSE is 0.005. Our MAE (mean absolute error) is 0.060. These measures show that this model is doing a good job at predicting responses for “new” data points. Additionally, the variance for these measures is relatively small, showing that these estimates are probably pretty close to the true predictive ability. This means that the MLR model can still be used to predict the COVID death rate.
Conclusion
- We employed the two automatic search procedures, criterion based approaches, and the LASSO technique to select a final model:
\(\hat{death\_rate}=-0.375+0.086*I(location=inland)+0.002*vac-0.004*labor+0.060*ln(tests)+0.026*ln(area)+0.093*ln(unemploy)+0.035*ln(bed)\) where vac is the vaccination rate, labor is the labor rate, unemploy is the unemployment rate, bed is the hospital available beds per person.
From this model, we can see that as the while the location is inland, unemployment, land area, and test participation increases, the predicted death rate increases. This is probably because inland counties, counties with high unemployment generally have poor economy so that poor medical conditions, which may finally lead to increased death rate; while the large size of the county may lead to increased death rate due to dense population, but still needs further investigation. In addition, the high death rate is from increased test participation is somewhat surprising, but it may be because more potential positive patients are able to be detected. However, the death rate also increases by vaccination rate increasing. The main suspicion is that the model may still miss some important information and more potential variables should be still needed.
Overall, thought there is suspision of underfitting, from our 10-fold cross validation, we see that our model still has good predictive ability for new points. However, this model was only built on data on the county level, so there may be some special information missing, but the overall model reflects the correlation and relationship between the death and some predictors in California.